Aptagen, LLC
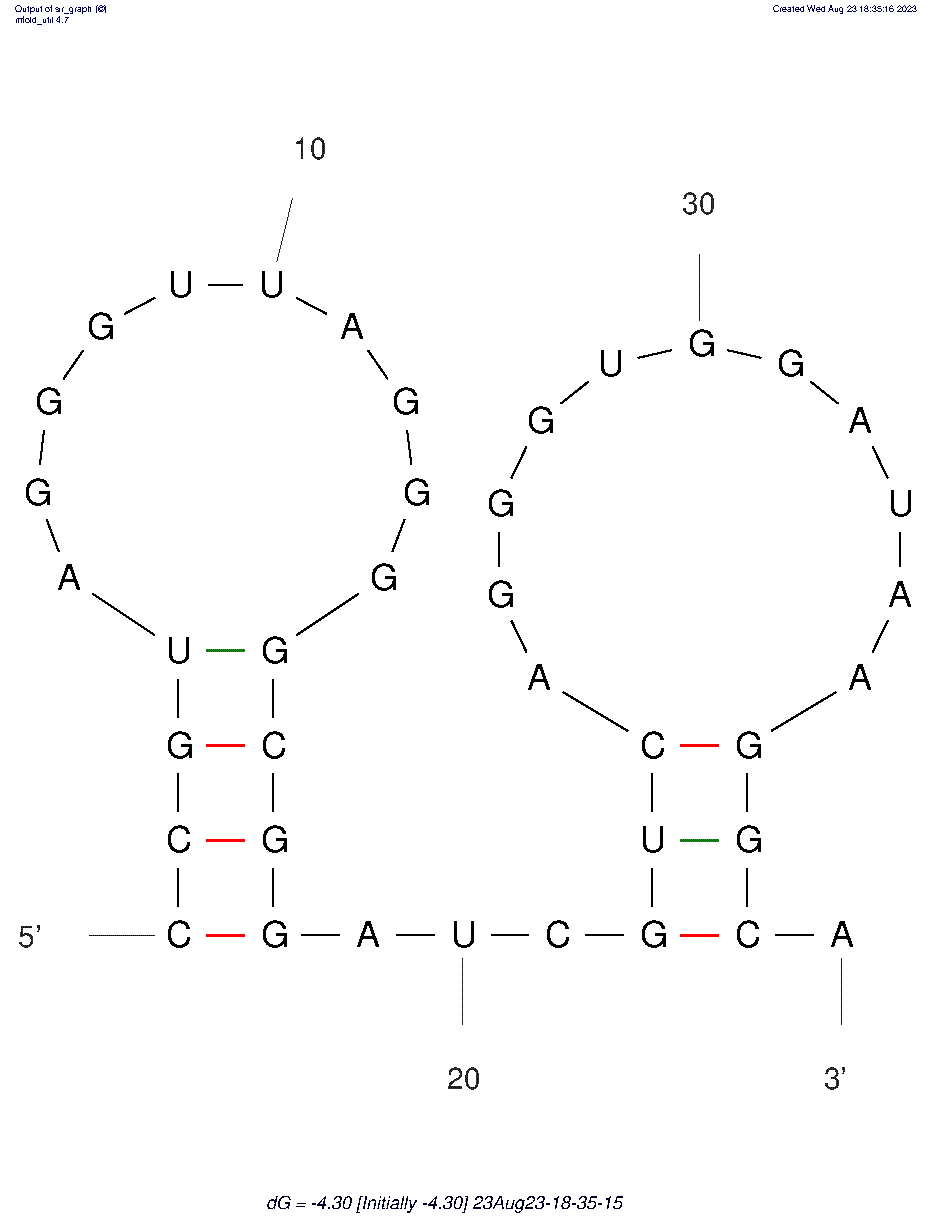
Crimean-Congo Hemorrhagic fever (ID# 8245)

RNA
Nucleoprotein (NP)
Protein
6.62x10^-10 nM (reported value)
In this study, an aptamer library consisting of 810 pmol was used in PBS as a binding buffer. The library was subjected to a heat treatment at 95°C for 5 minutes, followed by rapid cooling on an ice bath. The cooled library was then mixed with 70 μl of magnetic beads (MB) and incubated at 4°C with gentle shaking for 1 hour. After incubation, unbound aptamers were removed by precipitation, and the resulting supernatant was retained. This supernatant was combined with 70 μl of another set of magnetic beads (NP-MB) and incubated for 90 minutes at 4°C with gentle shaking. The mixture was then precipitated using a magnet, and the supernatant was discarded. The collected pellet was washed with 70 μl of PBS, followed by suspension in 75 μl of RNase DNase-free water at 70°C. A portion of 15 μl from the suspended pellet was stored at -20°C, while the remaining 60 μl was employed as a template in 4 PCR reactions for subsequent rounds of SELEX (Systematic Evolution of Ligands by Exponential enrichment). This process enables the selection and amplification of target-specific aptamers for further study.
4°C
NA If the oligo is a known aptamer sequence: For binding studies, perform a refolding protocol to ensure proper function (i.e. binding to antigen or target). Refer to the aptamer reference source for the appropriate refolding parameters and binding conditions. Note: it is unknown whether aptamer functions properly without refolding.
'
5'-dCpdCpdGpdTpdApdGpdGpdGpdTpdTpdApdGpdGpdGpdGpdCpdGpdGpdApdTpdCpdGpdTpdCpdApdGpdGpdGpdTpdGpdGpdApdTpdApdApdGpdGpdCpdAp-3'


39
2940.15
375
Note: Information on this aptamer oligo was obtained from the literature and hasn't been validated by Aptagen.
Jalali, T., Salehi-Vaziri, M., Pouriayevali, M. H., & Gargari, S. L. M. (2021). Aptamer based diagnosis of crimean-congo hemorrhagic fever from clinical specimens. Scientific Reports, 11(1). https://doi.org/10.1038/s41598-021-91826-8
Have your aptamer oligo synthesized ORDER



We are always looking for ways to improve. Please tell us what you think.