Aptagen, LLC
 DNA
Acute lymphoblastic leukemia (CCRF-CEM)
Cells
0.8 nM (reported value)
Binding buffer: 0.1mg/ml yeast tRNA and 1 mg/ml BSA into wash buffer (4.5g/L glucose, 5 mM MgCl2 in Dulbecco's PBS with CaCl2 and MgCl2)
4°C
NA If the oligo is a known aptamer sequence: For binding studies, perform a refolding protocol to ensure proper function (i.e. binding to antigen or target). Refer to the aptamer reference source for the appropriate refolding parameters and binding conditions. Note: it is unknown whether aptamer functions properly without refolding.
Recognizes and binds to cell lines Molt-4 (T cell Acute Lymphoblastic Leukemia [ALL]), Sup-T1 (T cell ALL), and Jurkat (T cell ALL).
This aptamer was found to bind to protein tyrosine kinase 7 (PTK7) (Shangguan 2008).
DNA
Acute lymphoblastic leukemia (CCRF-CEM)
Cells
0.8 nM (reported value)
Binding buffer: 0.1mg/ml yeast tRNA and 1 mg/ml BSA into wash buffer (4.5g/L glucose, 5 mM MgCl2 in Dulbecco's PBS with CaCl2 and MgCl2)
4°C
NA If the oligo is a known aptamer sequence: For binding studies, perform a refolding protocol to ensure proper function (i.e. binding to antigen or target). Refer to the aptamer reference source for the appropriate refolding parameters and binding conditions. Note: it is unknown whether aptamer functions properly without refolding.
Recognizes and binds to cell lines Molt-4 (T cell Acute Lymphoblastic Leukemia [ALL]), Sup-T1 (T cell ALL), and Jurkat (T cell ALL).
This aptamer was found to bind to protein tyrosine kinase 7 (PTK7) (Shangguan 2008).
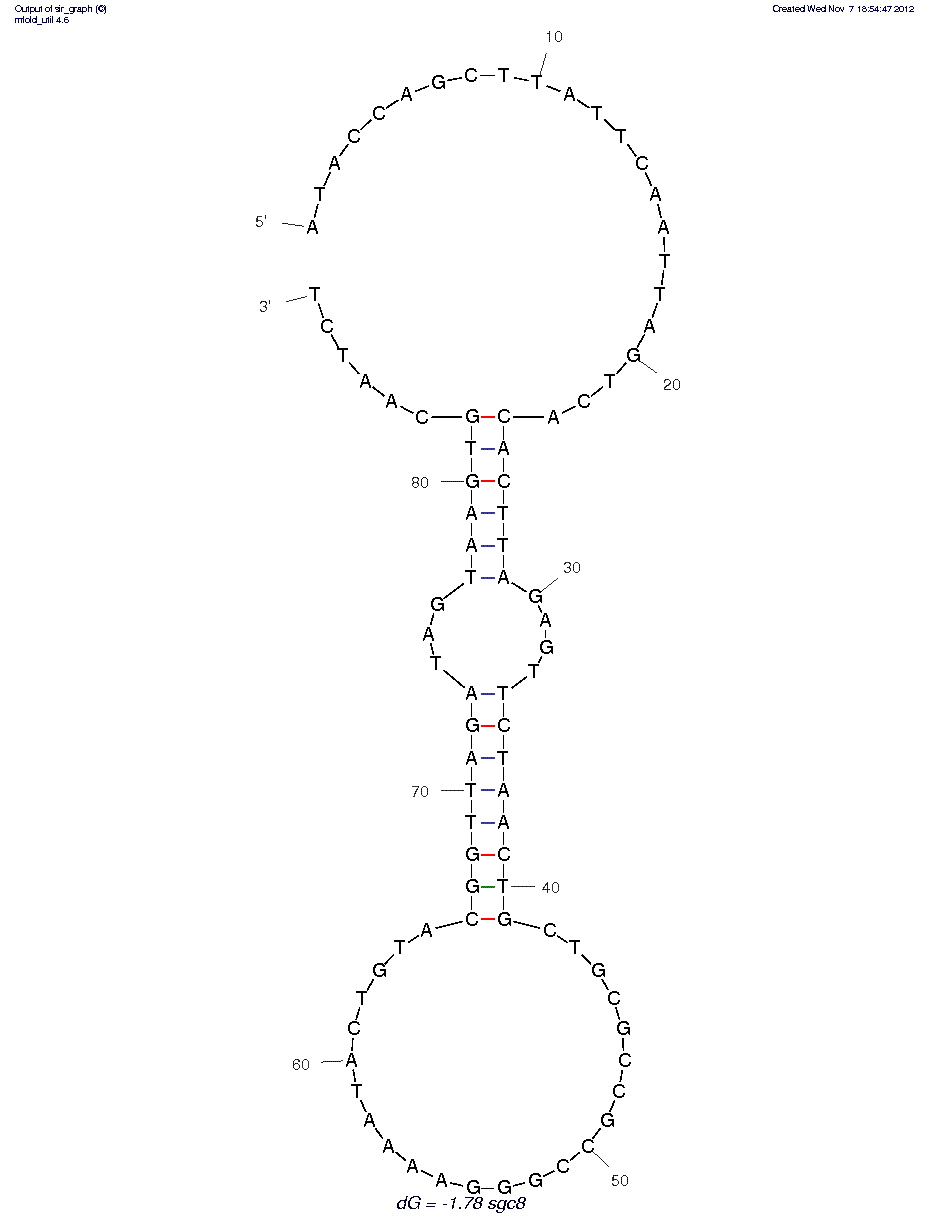
CCRF-CEM (sgc8) (ID# 63)

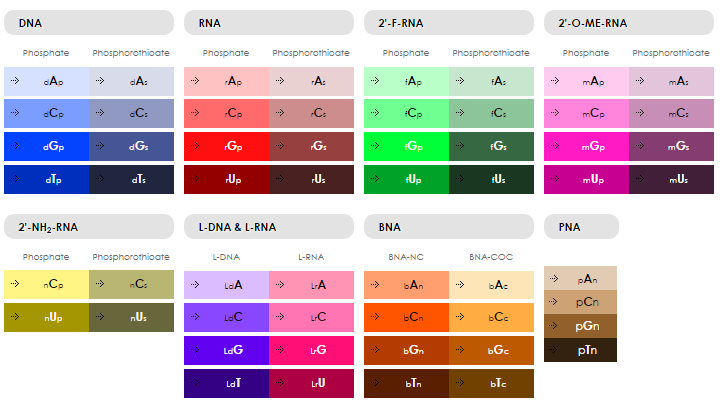
5'dApdTpdApdCpdCpdApdGpdCpdTpdTpdApdTpdTpdCpdApdApdTpdTpdApdGpdTpdCpdApdCpdApdCpdTpdTpdApdGpdApdGpdTpdTpdCpdTpdApdApdCpdTpdGpdCpdTpdGpdCpdGpdCpdCpdGpdCpdCpdGpdGpdGpdApdApdApdApdTpdApdCpdTpdGpdTpdApdCpdGpdGpdTpdTpdApdGpdApdTpdApdGpdTpdApdApdGpdTpdGpdCpdApdApdTpdCpdTp3'
88
27106.7 g/mole
859200 L/(mole·cm)
42.05%
1.16
31.55
Shangguan et al. ƒ??Aptamers evolved from live cells as effective molecular probes for cancer study.ƒ? Proceedings of the National Academy of Sciences of the United States of America, 103 (2006): 11838-43. Sequence was given in: Shangguan et al. "Optimization and Modifications of Aptamers Selected from Live Cancer Cell Lines." Chem. Biochem., 8, (2007): 603-6. Target protein identification:
Note: Information on this aptamer oligo was obtained from the literature and hasn't been validated by Aptagen.
Shangguan et al. ƒ??Aptamers evolved from live cells as effective molecular probes for cancer study.ƒ? Proceedings of the National Academy of Sciences of the United States of America, 103 (2006): 11838-43. Sequence was given in: Shangguan et al. "Optimization and Modifications of Aptamers Selected from Live Cancer Cell Lines." Chem. Biochem., 8, (2007): 603-6. Target protein identification:
D Shangguan et al. Cell-specific aptamer probes for membrane protein elucidation in cancer cells. J. Proteome Res. 7:2133-2139.
Have your aptamer oligo synthesized ORDER NOW




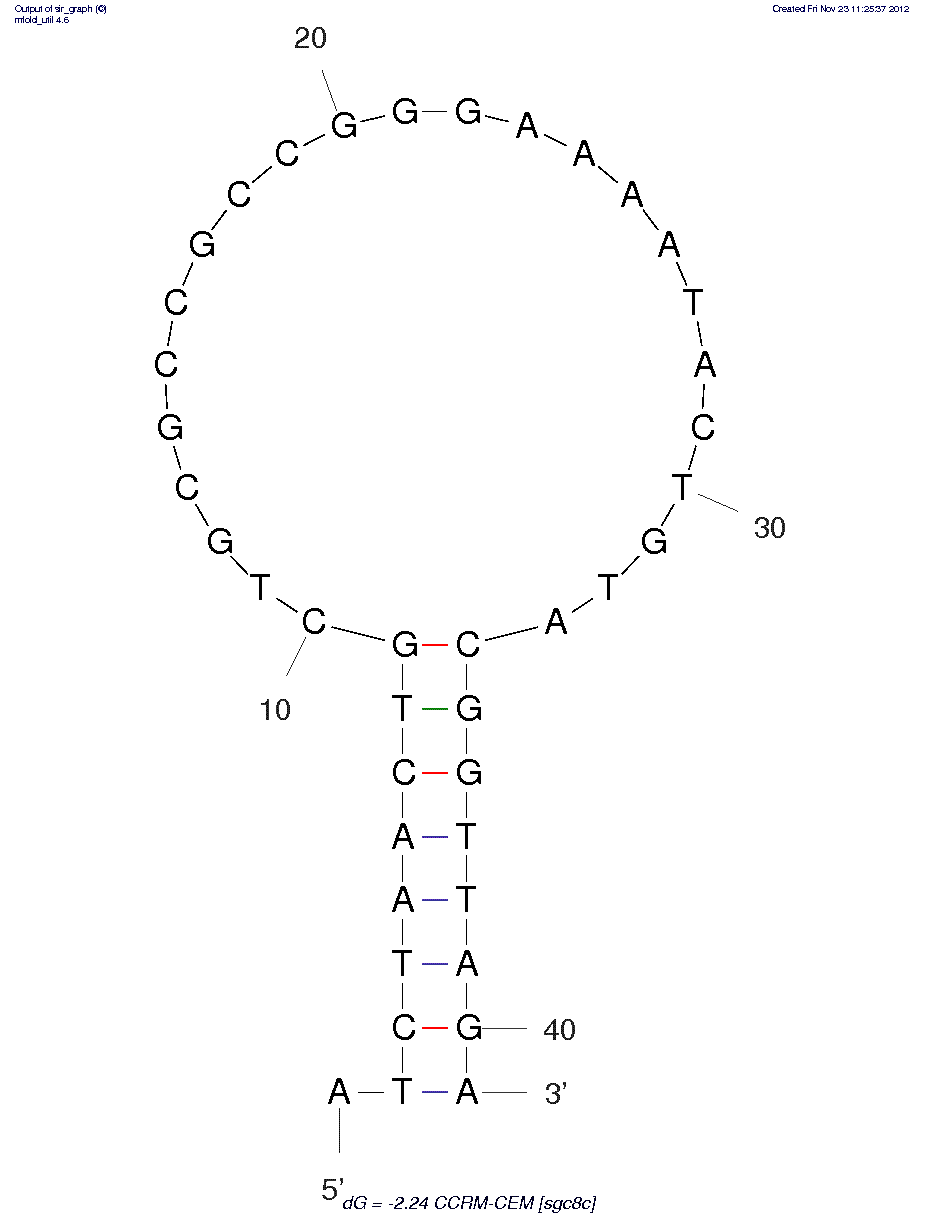
We are always looking for ways to improve. Please tell us what you think.